Clc genomics workbench gvcf12/7/2022  Based on sequence divergence of mtCOI (≥ 3.5% divergence), B. Although the use of ≥3.5% mtCOI divergence as the criterion for species delimitation has been occasionally shown to be inadequate, it has been widely used to differentiate species.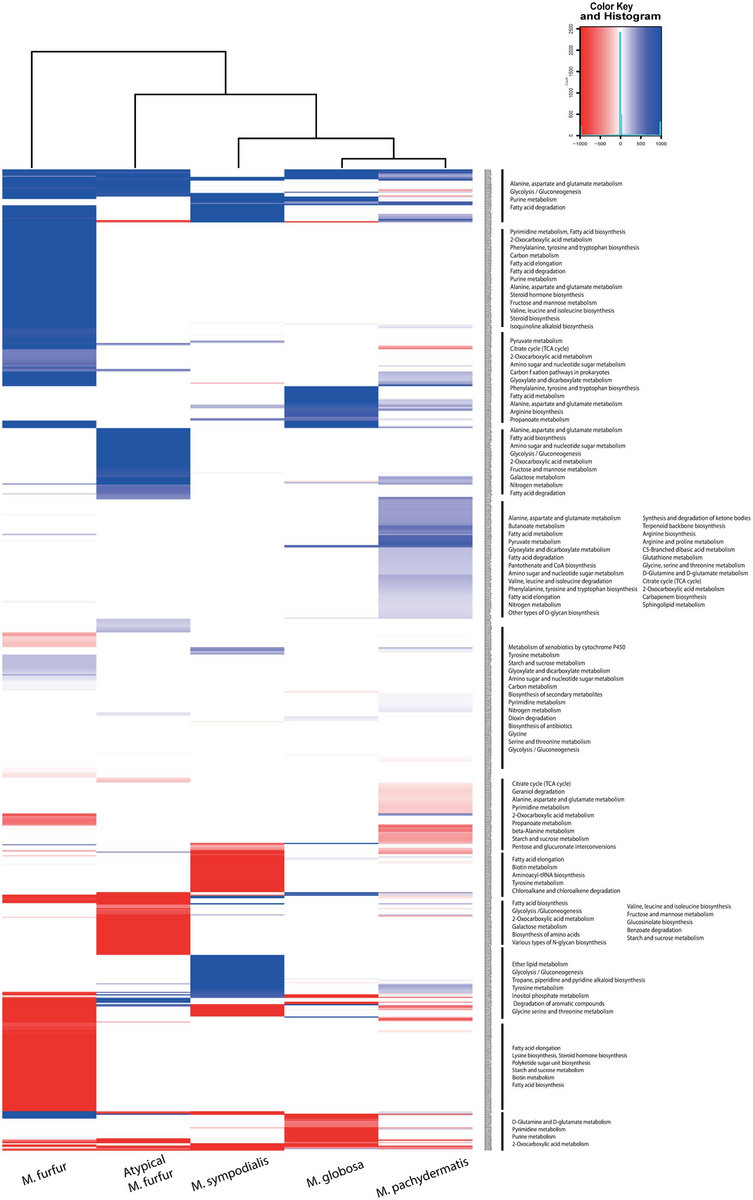 
These putative whitefly species differ in many biological aspects such as host range, resistance to insecticides, specificity and capacity of virus transmission and composition of harbored symbionts. tabaci) are now known to be as a cryptic species complex, based on recent molecular phylogenetic analyses and evidence of reproductive incompatibility. 
tabaci-transmitted viruses on a global scale include cotton, cassava, tomato, sweet potato, cucurbits and other crop plant species. These whiteflies can potentially vector over 300 plant viruses, mostly viruses in the genus Begomovirus. The whiteflies can infest as many as 1000 plant species and they damage host plants by infestation, but more importantly by transmitting plant viruses. The present study provides new genomic data and information resources for Asia II 1 that will not only contribute to the species delimitation of whitefly, but also help in conceiving future research studies to develop more targeted management strategies against whitefly.īemisia tabaci (Hemiptera: Aleyrodidae), commonly known as ‘whiteflies’ are phloem sap sucking pests some of which have become a major constraint to important food, fiber and ornamental crops worldwide. 
These genetic differences between Asia II 1 and MEAM1 may underlie the major biological differences between the two species such as virus transmission, insecticide resistance, and range of host plants. The functional analysis revealed that some of the genes are involved in virus transmission including 4 genes in Tomato yellow leaf curl virus (TYLCV) transmission, 96 in Tomato crinivirus (ToCV) transmission, and 14 genes in insecticide resistance. In Total, 1294 genes were detected with high impact variants. The sequence comparison with MEAM1 identified 2,327,972 SNPs and 202,479 INDELs. Sequencing of the nuclear genome of Asia II 1 with Illumina HiSeq and MiSeq generated 198.90 million reads that covers 88% of the reference genome. The purpose of this study is to find genomic differences between these two species. Asia II 1 is an indigenous species found on the Indian sub-continent and south-east Asia while the species named as Middle East Asia Minor 1 (MEAM1), likely originated from the Middle-East and has spread worldwide in recent decades. 
tabaci are now known to be a complex of cryptic species that differ from each other in many characteristics such as mode of interaction with viruses, invasiveness, and resistance to insecticides. Clc genomics workbench gvcf full#We support all the major next generation sequencing platforms, such as SOLiD, Ion Torrent, Complete Genomics, 454, Illumina Genome Analyzer and of course also Sanger, and we are working closely together with all the instrument vendors to ensure full integration in the ongoing development.ĬLC Genomics Workbench supports read mapping as well as de novo assembly of hybrid data.Whiteflies ( Bemisia tabaci) are phloem sap-sucking pests that because of their broad host range and ability to transmit viruses damage crop plants worldwide. It includes a number of features within the fields of genomics, transcriptomics and epigenomics, and additionally it includes all the tools of CLC Main Workbench. Clc genomics workbench gvcf mac os#We have overcome the challenge to analyze high-throughput sequencing data faster than it is produced by implementing a SIMD-accelerated assembly algorithm in our next generation sequencing solution, CLC Genomics Workbench – a cross-platform desktop application with a graphical user-interface.ĬLC Genomics Workbench, for analyzing and visualizing next generation sequencing data, incorporates cutting-edge technology and algorithms, while also supporting and integrating with the rest of your typical NGS workflow.ĬLC Genomics Workbench is available for Windows, Mac OS X, and Linux platforms. Dominating the high-throughput sequencing data analysis challenge 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
